Artificial Intelligence has fast taken up adoption across various industry domains. Whether it’s playing chess, predicting disease, forecasting the stock market rally, simulating the betting odds on sports games, detecting credit card fraud, creating new artworks or composing Shakespeare’s plays, the applications of Artificial Intelligence have continued to grow over the last decade.
DeepMind is a London-based AI company owned by Google that creates neural networks that learn to play video games. The company is known for the Neural Turing machine that can copy, sort, and recall simple machine algorithms.
However, until last year, DeepMind was not a name well-known in structural biology. Yet, Deep Mind’s creative recombination has enabled it to take its learnings from other industry domains and mesh them with its talent and expertise in machine learning and neural networks to solve one of structural biology’s most challenging problems – The Protein Folding Problem.
EMBL (European Bioinformatics Institute – EMBL EBI) and DeepMind profiled the proteins across the animal, fungi, and bacteria kingdoms to solve the Protein Folding problem. Leveraging domain knowledge in proteomics and technical machine learning expertise, DeepMind and EMBL crafted artificial neural networks to profile millions of proteins and map their structure.
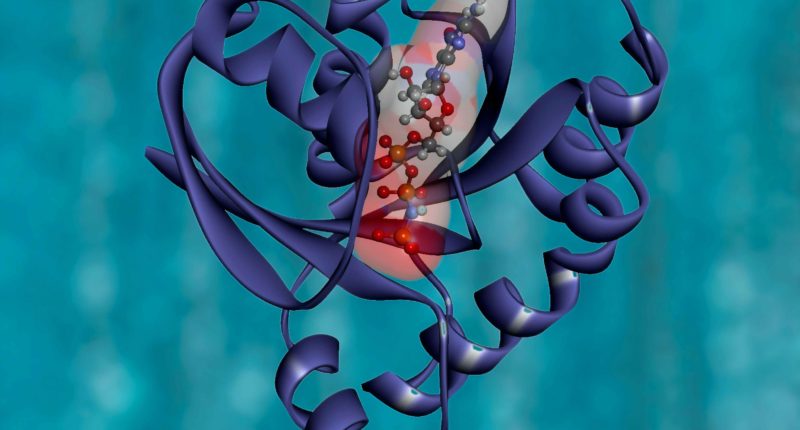
Proteins are the building blocks of life. All living animal cells are built of proteins. A key signature of a protein is its unique 3-D structure. The 3-D structure or the protein’s folding pattern indicates these proteins’ function and behavior. The folding process for proteins can be highly complex, and the laws of physics guide the folding process and patterns.
Since the 1950s, biologists, pharmacists, chemists, and medical and drug researchers have continued to understand how proteins fold and what the protein folding mechanism is. Researchers have also grappled with mapping the native structure of the protein. In the realm of science, the Protein Folding Challenge is an optimization problem that currently requires long hours in the lab, leveraging thorough trial and testing with X-ray crystallography to best determine a protein’s folding structure and specificity. The protein folding problem is complex as millions of proteins exist in the animal kingdom, and many have unique folding structures. Imagine trying to predict all their structures.
DeepMind’s AlphaFold calculated the structure of over 200 million proteins using neural networks. The company’s protein folding data model has significant precision and high accuracy compared to gold standard tests like X-ray crystallography. Most Alphafold predictions had an accuracy of 90% on highly complex derived proteins. The data from AlphaFold is available in an open UniProt database, freely available for researchers, academics, and scientists worldwide.
Given that the Protein Folding Problem has been resolved in medical sciences, the insights from protein structure will help accelerate society’s understanding of how cells operate, how biological cells communicate, and how enzymes and disease interact.
Referring to AlphaFold and EMBL-EBI’s protein folding discovery, Dr. Eric Topol, Founder and Director of the Scripps Research Translational Institute, commented, “Determining the 3D structure of a single protein used to take many months or years. It now takes seconds. AlphaFold has already accelerated and enabled massive discoveries, including cracking the structure of the nuclear pore complex. And with this new addition of structure illuminating nearly the entire protein universe, we can expect more biological mysteries to be solved daily.”
Alphafold’s advances continue to revolutionize science, and perhaps its technology on proteins will help enable new drug targets for Cancer, HIV or Alzheimer’s or even discover therapies for chronic illnesses like diabetes, heart disease or high blood pressure.



Comments are closed.